Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Immune Response after Anti-SARS-CoV-2 mRNA Vaccination in Relation to Cellular Immunity, Vitamin D and Comorbidities in Hemodialysis Patients
Microorganisms 2024, 12(5), 861; https://doi.org/10.3390/microorganisms12050861 (registering DOI) - 25 Apr 2024
Abstract
In the global threat of SARS-CoV-2, individuals undergoing maintenance dialysis represent a vulnerable population with an increased risk of severe COVID-19 outcomes. Therefore, immunization against SARS-CoV-2 is an essential component of healthcare strategy for these patients. Existing data indicate that they tend to
[...] Read more.
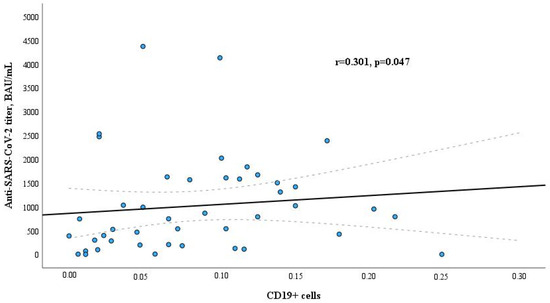
In the global threat of SARS-CoV-2, individuals undergoing maintenance dialysis represent a vulnerable population with an increased risk of severe COVID-19 outcomes. Therefore, immunization against SARS-CoV-2 is an essential component of healthcare strategy for these patients. Existing data indicate that they tend to exhibit a reduced immune response to vaccines compared to the general population. Our study aimed to assess both humoral and cellular immune responses following two doses of an anti-SARS-CoV-2 mRNA vaccine, an ability to maintain adequate antibody titers over time, and potential relations with vitamin D, comorbidities and other factors in hemodialysis patients based on a single center experience. A total of 41/45 patients (91.1%) responded to the second dose of the anti-SARS-CoV-2 mRNA vaccine. The titer of anti-SARS-CoV-2 IgG class antibodies and levels of T cells three to four weeks after vaccination were lower in dialysis patients than in healthy controls. Antibodies titer in dialysis patients had a positive correlation with B lymphocytes and was related to cardiovascular diseases. The level of CD4+ cells had a negative correlation with hemodialysis vintage, as did the vitamin D level with post-vaccination seroconversion and decline in anti-SARS-CoV-2 antibodies titer during six months after vaccination. Hemodialysis patients had decreased amounts of CD4+ and CD8+ cells and lower levels of anti-SARS-CoV-2 antibodies than healthy controls. Therefore, chronic hemodialysis could lead to diminished cellular immunity and humoral immune response to the anti-SARS-CoV-2 mRNA vaccination and reduced protection from COVID-19. Comorbidity in cardiovascular diseases was associated with a lower level of specific anti-SARS-CoV-2 antibody titer. Vitamin D may be important in maintaining stable levels of anti-SARS-CoV-2 antibodies, while the duration of dialysis treatment could be one of the factors decreasing anti-SARS-CoV-2 antibody titer and determining lower CD4+ cell counts.
Full article
(This article belongs to the Special Issue Immune Modulation to SARS-CoV-2 Vaccination and Infection)
►
Show Figures
Open AccessArticle
Multi-Omics of Campylobacter jejuni Growth in Chicken Exudate Reveals Molecular Remodelling Associated with Altered Virulence and Survival Phenotypes
by
Lok Man, Pamela X. Y. Soh, Tess E. McEnearney, Joel A. Cain, Ashleigh L. Dale and Stuart J. Cordwell
Microorganisms 2024, 12(5), 860; https://doi.org/10.3390/microorganisms12050860 (registering DOI) - 25 Apr 2024
Abstract
Campylobacter jejuni is the leading cause of foodborne human gastroenteritis in the developed world. Infections are largely acquired from poultry produced for human consumption and poor food handling is thus a major risk factor. Chicken exudate (CE) is a liquid produced from defrosted
[...] Read more.
Campylobacter jejuni is the leading cause of foodborne human gastroenteritis in the developed world. Infections are largely acquired from poultry produced for human consumption and poor food handling is thus a major risk factor. Chicken exudate (CE) is a liquid produced from defrosted commercial chicken products that facilitates C. jejuni growth. We examined the response of C. jejuni to growth in CE using a multi-omics approach. Changes in the C. jejuni proteome were assessed by label-based liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS). We quantified 1328 and 1304 proteins, respectively, in experiments comparing 5% CE in Mueller–Hinton (MH) medium and 100% CE with MH-only controls. These proteins represent 81.8% and 80.3% of the predicted C. jejuni NCTC11168 proteome. Growth in CE induced profound remodelling of the proteome. These changes were typically conserved between 5% and 100% CE, with a greater magnitude of change observed in 100% CE. We confirmed that CE induced C. jejuni biofilm formation, as well as increasing motility and resistance against oxidative stress, consistent with changes to proteins representing those functions. Assessment of the C. jejuni metabolome showed CE also led to increased intracellular abundances of serine, proline, and lactate that were correlated with the elevated abundances of their respective transporters. Analysis of carbon source uptake showed prolonged culture supernatant retention of proline and succinate in CE-supplemented medium. Metabolomics data provided preliminary evidence for the uptake of chicken-meat-associated dipeptides. C. jejuni exposed to CE showed increased resistance to several antibiotics, including polymyxin B, consistent with changes to tripartite efflux system proteins and those involved in the synthesis of lipid A. The C. jejuni CE proteome was also characterised by very large increases in proteins associated with iron acquisition, while a decrease in proteins containing iron–sulphur clusters was also observed. Our data suggest CE is both oxygen- and iron-limiting and provide evidence of factors required for phenotypic remodelling to enable C. jejuni survival on poultry products.
Full article
(This article belongs to the Special Issue Foodborne Bacteria–Host Interactions 2.0)
►▼
Show Figures
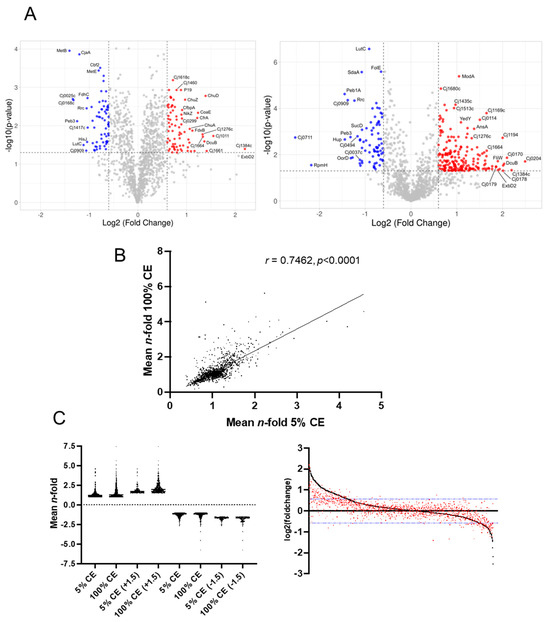
Figure 1
Open AccessEditorial
Editorial: Impact of Special Issue “The Microbial Population of the Gastrointestinal Tract of Animals: Impacts on Host Physiology”
by
Jeferson M. Lourenco and Todd R. Callaway
Microorganisms 2024, 12(5), 859; https://doi.org/10.3390/microorganisms12050859 (registering DOI) - 25 Apr 2024
Abstract
In recent years, there has been an exponential increase in the number of papers that have investigated the microbiome of animals and humans [...]
Full article
(This article belongs to the Special Issue The Microbial Population of the Gastrointestinal Tract of Animals: Impacts on Host Physiology)
Open AccessArticle
Temporal Dynamics of Fungal Communities in Alkali-Treated Round Bamboo Deterioration under Natural Weathering
by
Shuaibo Han, Xiaojiao An, Xiaolong He, Xin Ren, John Sichone, Xinxing Wu, Yan Zhang, Hui Wang and Fangli Sun
Microorganisms 2024, 12(5), 858; https://doi.org/10.3390/microorganisms12050858 (registering DOI) - 25 Apr 2024
Abstract
Microbes naturally inhabit bamboo-based materials in outdoor environments, sequentially contributing to their deterioration. Fungi play a significant role in deterioration, especially in environments with abundant water and favorable temperatures. Alkali treatment is often employed in the pretreatment of round bamboo to change its
[...] Read more.
Microbes naturally inhabit bamboo-based materials in outdoor environments, sequentially contributing to their deterioration. Fungi play a significant role in deterioration, especially in environments with abundant water and favorable temperatures. Alkali treatment is often employed in the pretreatment of round bamboo to change its natural elastic and aesthetic behaviors. However, little research has investigated the structure and dynamics of fungal communities on alkali-treated round bamboo during natural deterioration. In this work, high-throughput sequencing and multiple characterization methods were used to disclose the fungal community succession and characteristic alterations of alkali-treated round bamboo in both roofed and unroofed habitats throughout a 13-week deterioration period. In total, 192 fungal amplicon sequence variants (ASVs) from six phyla were identified. The fungal community richness of roofed bamboo samples declined, whereas that of unroofed bamboo samples increased during deterioration. The phyla Ascomycota and Basidiomycota exhibited dominance during the entire deterioration process in two distinct environments, and the relative abundance of them combined was more than 99%. A distinct shift in fungal communities from Basidiomycota dominant in the early stage to Ascomycota dominant in the late stage was observed, which may be attributed to the increase of moisture and temperature during succession and the effect of alkali treatment. Among all environmental factors, temperature contributed most to the variation in the fungal community. The surface of round bamboo underwent continuous destruction from fungi and environmental factors. The total amount of cell wall components in bamboo epidermis in both roofed and unroofed conditions presented a descending trend. The content of hemicellulose declined sharply by 8.3% and 11.1% under roofed and unroofed environments after 9 weeks of deterioration. In addition, the contact angle was reduced throughout the deterioration process in both roofed and unroofed samples, which might be attributed to wax layer removal and lignin degradation. This study provides theoretical support for the protection of round bamboo under natural weathering.
Full article
(This article belongs to the Section Plant Microbe Interactions)
►▼
Show Figures
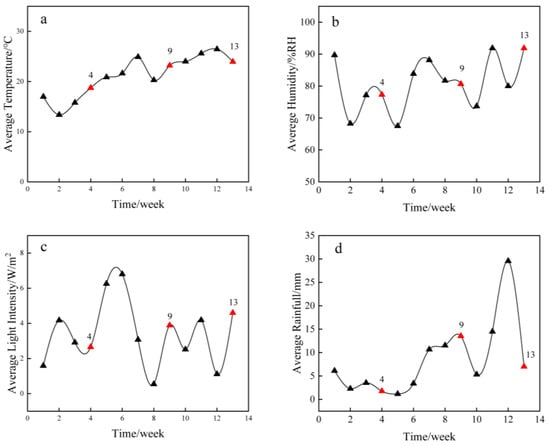
Figure 1
Open AccessCase Report
From Investigating a Case of Cellulitis to Exploring Nosocomial Infection Control of ST1 Legionella pneumophila Using Genomic Approaches
by
Charlotte Michel, Fedoua Echahidi, Sammy Place, Lorenzo Filippin, Vincent Colombie, Nicolas Yin, Delphine Martiny, Olivier Vandenberg, Denis Piérard and Marie Hallin
Microorganisms 2024, 12(5), 857; https://doi.org/10.3390/microorganisms12050857 (registering DOI) - 25 Apr 2024
Abstract
Legionella pneumophila can cause a large panel of symptoms besides the classic pneumonia presentation. Here we present a case of fatal nosocomial cellulitis in an immunocompromised patient followed, a year later, by a second case of Legionnaires’ disease in the same ward. While
[...] Read more.
Legionella pneumophila can cause a large panel of symptoms besides the classic pneumonia presentation. Here we present a case of fatal nosocomial cellulitis in an immunocompromised patient followed, a year later, by a second case of Legionnaires’ disease in the same ward. While the first case was easily assumed as nosocomial based on the date of symptom onset, the second case required clear typing results to be assigned either as nosocomial and related to the same environmental source as the first case, or community acquired. To untangle this specific question, we applied core-genome multilocus typing (MLST), whole-genome single nucleotide polymorphism and whole-genome MLST methods to a collection of 36 Belgian and 41 international sequence-type 1 (ST1) isolates using both thresholds recommended in the literature and tailored threshold based on local epidemiological data. Based on the thresholds applied to cluster isolates together, the three methods gave different results and no firm conclusion about the nosocomial setting of the second case could been drawn. Our data highlight that despite promising results in the study of outbreaks and for large-scale epidemiological investigations, next-generation sequencing typing methods applied to ST1 outbreak investigation still need standardization regarding both wet-lab protocols and bioinformatics. A deeper evaluation of the L. pneumophila evolutionary clock is also required to increase our understanding of genomic differences between isolates sampled during a clinical infection and in the environment.
Full article
(This article belongs to the Special Issue Clinical and Environmental Surveillance for the Prevention of Legionellosis 2.0)
Open AccessCommunication
Trichinella spiralis Infecting Wild Boars in West, Southwest, and Northwest of Romania: Evidence of an Underrated Risk
by
Ana-Maria Marin, Tudor Rareș Olariu, Dan-Cornel Popovici, Gianluca Marucci, Sorin Morariu, Daian Popa and Narcisa Mederle
Microorganisms 2024, 12(5), 856; https://doi.org/10.3390/microorganisms12050856 (registering DOI) - 25 Apr 2024
Abstract
The species of the genus Trichinella are etiological agents distributed all over the world and are able to infect mammals, birds, and reptiles. Trichinella spiralis is the species most adapted to domestic and wild pigs and is also the most important etiological agent
[...] Read more.
The species of the genus Trichinella are etiological agents distributed all over the world and are able to infect mammals, birds, and reptiles. Trichinella spiralis is the species most adapted to domestic and wild pigs and is also the most important etiological agent of trichinellosis. The wild boar (Sus scrofa) is a nocturnal omnivorous mammal belonging to the Suidae family. S. scrofa has a great appetite and its diet includes a variety of small prey such as mice, rats, and other rodents, as well as carcasses of larger animals. The aim of this study was the identification and the molecular characterization of Trichinella larvae isolated from the muscle tissue of S. scrofa specimens collected in different counties of Romania. The muscle samples were examined by artificial digestion and the larvae identified at the species level by multiplex PCR. T. spiralis, a species that is able to infect a considerable number of different host species including humans, was identified. In Romania, S. scrofa is an important reservoir species for T. spiralis and plays an important role in linking the domestic and the wild cycle of Trichinella, with serious repercussions for human health.
Full article
(This article belongs to the Section Parasitology)
Open AccessReview
Interaction of Trypanosoma cruzi, Triatomines and the Microbiota of the Vectors—A Review
by
Günter A. Schaub
Microorganisms 2024, 12(5), 855; https://doi.org/10.3390/microorganisms12050855 (registering DOI) - 25 Apr 2024
Abstract
This review summarizes the interactions between Trypanosoma cruzi, the etiologic agent of Chagas disease, its vectors, triatomines, and the diverse intestinal microbiota of triatomines, which includes mutualistic symbionts, and highlights open questions. T. cruzi strains show great biological heterogeneity in their development
[...] Read more.
This review summarizes the interactions between Trypanosoma cruzi, the etiologic agent of Chagas disease, its vectors, triatomines, and the diverse intestinal microbiota of triatomines, which includes mutualistic symbionts, and highlights open questions. T. cruzi strains show great biological heterogeneity in their development and their interactions. Triatomines differ from other important vectors of diseases in their ontogeny and the enzymes used to digest blood. Many different bacteria colonize the intestinal tract of triatomines, but only Actinomycetales have been identified as mutualistic symbionts. Effects of the vector on T. cruzi are indicated by differences in the ability of T. cruzi to establish in the triatomines and in colonization peculiarities, i.e., proliferation mainly in the posterior midgut and rectum and preferential transformation into infectious metacyclic trypomastigotes in the rectum. In addition, certain forms of T. cruzi develop after feeding and during starvation of triatomines. Negative effects of T. cruzi on the triatomine vectors appear to be particularly evident when the triatomines are stressed and depend on the T. cruzi strain. Effects on the intestinal immunity of the triatomines are induced by ingested blood-stage trypomastigotes of T. cruzi and affect the populations of many non-symbiotic intestinal bacteria, but not all and not the mutualistic symbionts. After the knockdown of antimicrobial peptides, the number of non-symbiotic bacteria increases and the number of T. cruzi decreases. Presumably, in long-term infections, intestinal immunity is suppressed, which supports the growth of specific bacteria, depending on the strain of T. cruzi. These interactions may provide an approach to disrupt T. cruzi transmission.
Full article
(This article belongs to the Section Parasitology)
►▼
Show Figures
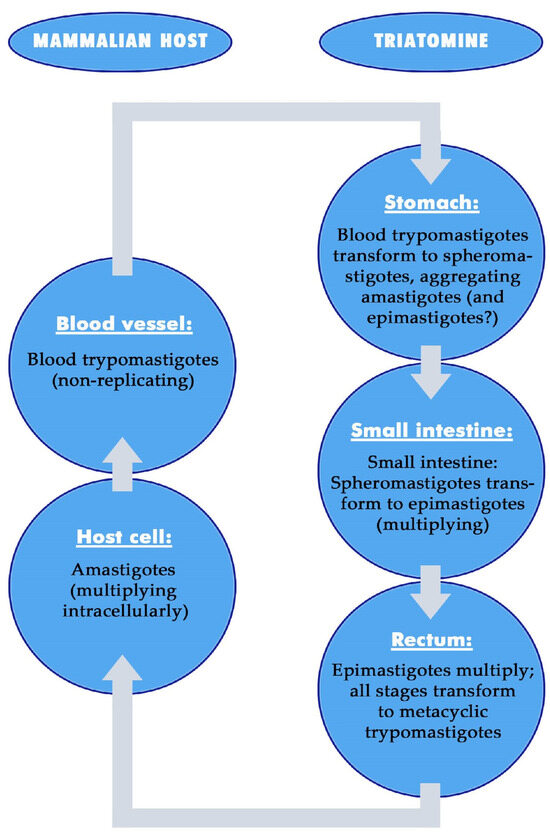
Figure 1
Open AccessReview
The Impact of Artificial Restoration of Alpine Grasslands in the Qilian Mountains on Vegetation, Soil Bacteria, and Soil Fungal Community Diversity
by
Xiaomei Yang, Qi Feng, Meng Zhu, Jutao Zhang, Linshan Yang and Ruolin Li
Microorganisms 2024, 12(5), 854; https://doi.org/10.3390/microorganisms12050854 (registering DOI) - 25 Apr 2024
Abstract
To understand how the soil microbial community structure responds to vegetation restoration in alpine mining areas, this study specifically examines the grassland ecosystem in the Qianmalong mining area of the Qilian Mountains after five years of artificial restoration. High-throughput sequencing methods were employed
[...] Read more.
To understand how the soil microbial community structure responds to vegetation restoration in alpine mining areas, this study specifically examines the grassland ecosystem in the Qianmalong mining area of the Qilian Mountains after five years of artificial restoration. High-throughput sequencing methods were employed to analyze soil bacteria and fungi microbial characteristics in diverse grassland communities. Combined with modifications in vegetation diversity as well as soil physicochemical properties, the impact of vegetation restoration on soil microbiome diversity in this alpine mining area was investigated. The findings indicated that the dominant plants were Cyperus rotundus, Carex spp., and Elymus nutans. As the extent of the grassland’s restoration increased, the number of plant species, importance values, and plant community diversity showed an increasing trend. The plant functional groups were mainly dominated by Cyperaceae, followed by Poaceae. Plant height, density, plant cover, frequency, and aboveground biomass showed an increasing trend, and soil water content (SWC) increased. While soil pH and soil electrical conductivity (EC) exhibited a declining trend, available phosphorus (AP), total phosphorus (TP), total nitrogen (TN), nitrate nitrogen (NO3-N), soil organic carbon (SOC), and soil water content (SWC) showed an increasing trend. The dominant bacterial communities were Actinobacteriota, Proteobacteria, Acidobacteriota, Chloroflexi, Firmicutes, and Gemmatimonadota, while the dominant fungal communities were Ascomycota, Mortierellomycota, Basidiomycota, unclassified_k_Fungi, and Glomeromycota. Significant differences were detected within soil microbial community composition among different degrees of restoration grasslands, with bacteria generally dominating over fungi. SWC, TP, and TN were found to be the main soil physicochemical factors affecting the distribution of soil bacterial communities’ structure; however, SOC, TN, and NO3-N were the primary factors influencing the soil distribution of fungal communities. The results of this study indicate that different degrees of vegetation restoration in alpine mining areas can significantly affect soil bacterial and fungal communities, and the degree of restoration has varying effects on the soil bacteria and fungi community structure in alpine mining areas.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures
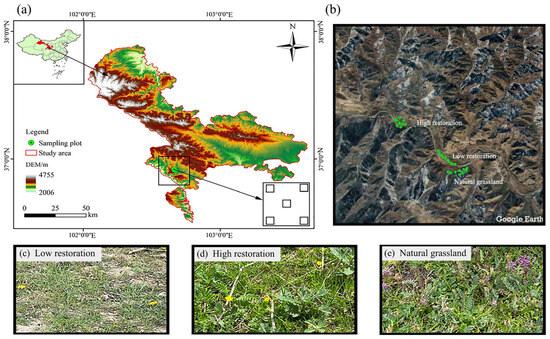
Figure 1
Open AccessArticle
Fodinisporobacter ferrooxydans gen. nov., sp. nov.—A Spore-Forming Ferrous-Oxidizing Bacterium Isolated from a Polymetallic Mine
by
Zhen Jiang, Xiutong Li, Zonglin Liang, Zebao Tan, Nan Zhou, Ying Liu, Zhenghua Liu, Huaqun Yin, Kun Luo, Supawadee Ingsriswang, Shuangjiang Liu and Chengying Jiang
Microorganisms 2024, 12(5), 853; https://doi.org/10.3390/microorganisms12050853 (registering DOI) - 25 Apr 2024
Abstract
A novel acidophilic, aerobic bacterium strain, MYW30-H2T, was isolated from a heap of polymetallic mine. Cells of strain MYW30-H2T were Gram-stain-positive, endospore-forming, motile, and rod-shaped. Strain MYW30-H2T grew at a temperature range of 30–45 °C (optimum 40 °C) and
[...] Read more.
A novel acidophilic, aerobic bacterium strain, MYW30-H2T, was isolated from a heap of polymetallic mine. Cells of strain MYW30-H2T were Gram-stain-positive, endospore-forming, motile, and rod-shaped. Strain MYW30-H2T grew at a temperature range of 30–45 °C (optimum 40 °C) and a pH range of 3.5–6.0 (optimum 4.0) in the presence of 0–0.5% (w/v) NaCl. Strain MYW30-H2T could grow heterotrophically on yeast extract and glucose, and grow mixotrophically using ferrous iron as an electron donor with yeast extract. Menaquinone-7 (MK-7) was the sole respiratory quinone of the strain. Iso-C15:0 and anteiso-C15:0 were the major cellular fatty acids. The 16S rRNA gene sequence analysis showed that MYW30-H2T was phylogenetically affiliated with the family Alicyclobacillaceae, and the sequence similarity with other Alicyclobacillaceae genera species was below 91.51%. The average amino acid identity value of the strain with its phylogenetically related species was 52.3–62.1%, which fell into the genus boundary range. The DNA G+C content of the strain was 44.2%. Based on physiological and phylogenetic analyses, strain MYW30-H2T represents a novel species of a new genus of the family Alicyclobacillaceae, for which the name Fodinisporobacter ferrooxydans gen. nov., sp. nov. is proposed. The type strain is MYW30-H2T (=CGMCC 1.17422T = KCTC 43278T).
Full article
(This article belongs to the Special Issue New Insights into the Diversity and Characterization of Extremophiles)
►▼
Show Figures
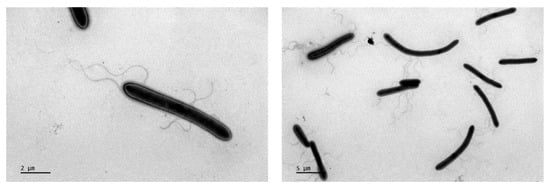
Figure 1
Open AccessReview
Developing the Common Marmoset as a Translational Geroscience Model to Study the Microbiome and Healthy Aging
by
Kelly R. Reveles, Alexana J. Hickmott, Kelsey A. Strey, Aaryn C. Mustoe, Juan Pablo Arroyo, Michael L. Power, Benjamin J. Ridenhour, Katherine R. Amato and Corinna N. Ross
Microorganisms 2024, 12(5), 852; https://doi.org/10.3390/microorganisms12050852 (registering DOI) - 25 Apr 2024
Abstract
Emerging data support associations between the depletion of the healthy gut microbiome and aging-related physiological decline and disease. In humans, fecal microbiota transplantation (FMT) has been used successfully to restore gut microbiome structure and function and to treat C. difficile infections, but its
[...] Read more.
Emerging data support associations between the depletion of the healthy gut microbiome and aging-related physiological decline and disease. In humans, fecal microbiota transplantation (FMT) has been used successfully to restore gut microbiome structure and function and to treat C. difficile infections, but its application to healthy aging has been scarcely investigated. The marmoset is an excellent model for evaluating microbiome-mediated changes with age and interventional treatments due to their relatively shorter lifespan and many social, behavioral, and physiological functions that mimic human aging. Prior work indicates that FMT is safe in marmosets and may successfully mediate gut microbiome function and host health. This narrative review (1) provides an overview of the rationale for FMT to support healthy aging using the marmoset as a translational geroscience model, (2) summarizes the prior use of FMT in marmosets, (3) outlines a protocol synthesized from prior literature for studying FMT in aging marmosets, and (4) describes limitations, knowledge gaps, and future research needs in this field.
Full article
(This article belongs to the Special Issue Fecal Microbiota Transplantation in Animals)
Open AccessReview
Advances in Gut Microbiota-Targeted Therapeutics for Metabolic Syndrome
by
Yu Gao, Wujuan Li, Xiaoyu Huang, Yuhong Lyu and Changwu Yue
Microorganisms 2024, 12(5), 851; https://doi.org/10.3390/microorganisms12050851 (registering DOI) - 24 Apr 2024
Abstract
Previous investigations have illuminated the significant association between the gut microbiome and a broad spectrum of health conditions, including obesity, diabetes, cardiovascular diseases, and psychiatric disorders. Evidence from certain studies suggests that dysbiosis of the gut microbiota may play a role in the
[...] Read more.
Previous investigations have illuminated the significant association between the gut microbiome and a broad spectrum of health conditions, including obesity, diabetes, cardiovascular diseases, and psychiatric disorders. Evidence from certain studies suggests that dysbiosis of the gut microbiota may play a role in the etiology of obesity and diabetes. Moreover, it is acknowledged that dietary habits, pharmacological interventions, psychological stress, and other exogenous factors can substantially influence the gut microbial composition. For instance, a diet rich in fiber has been demonstrated to increase the population of beneficial bacteria, whereas the consumption of antibiotics can reduce these advantageous microbial communities. In light of the established correlation between the gut microbiome and various pathologies, strategically altering the gut microbial profile represents an emerging therapeutic approach. This can be accomplished through the administration of probiotics or prebiotics, which aim to refine the gut microbiota and, consequently, mitigate the manifestations of associated diseases. The present manuscript evaluates the recent literature on the relationship between gut microbiota and metabolic syndrome published over the past three years and anticipates future directions in this evolving field.
Full article
(This article belongs to the Section Gut Microbiota)
►▼
Show Figures
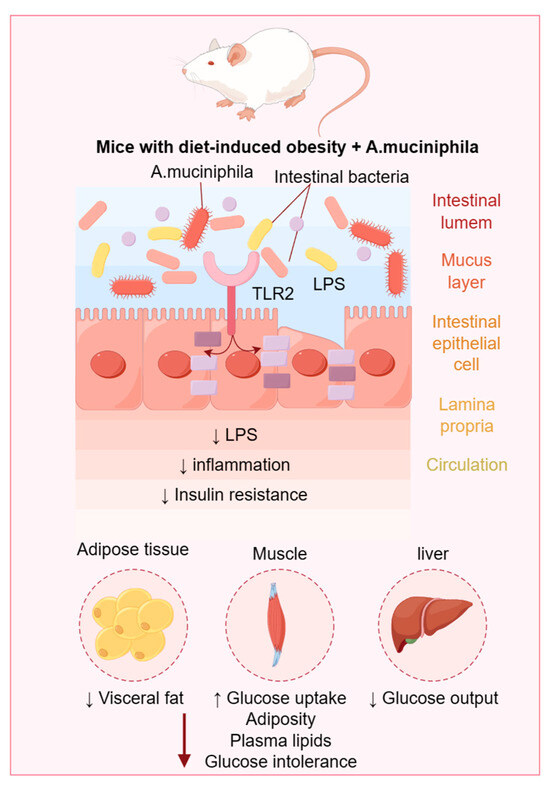
Figure 1
Open AccessArticle
Type I Cystatin Derived from Cysticercus pisiformis—Stefins, Suppresses LPS-Mediated Inflammatory Response in RAW264.7 Cells
by
Qianqian Yang, Jia Li, Lilan Zhang, Ningning Zhao, Xiaolin Sun and Zexiang Wang
Microorganisms 2024, 12(5), 850; https://doi.org/10.3390/microorganisms12050850 - 24 Apr 2024
Abstract
Cysticercus pisiformis is a kind of tapeworm larvae of Taenia pisiformis, which parasitizes the liver envelope, omentum, mesentery, and rectum of rodents such as rabbits. Cysteine protease inhibitors derived from helminth were immunoregulatory molecules of intermediate hosts and had an immunomodulatory function
[...] Read more.
Cysticercus pisiformis is a kind of tapeworm larvae of Taenia pisiformis, which parasitizes the liver envelope, omentum, mesentery, and rectum of rodents such as rabbits. Cysteine protease inhibitors derived from helminth were immunoregulatory molecules of intermediate hosts and had an immunomodulatory function that regulates the production of inflammatory factors. Thus, in the present research, the recombinant Stefin of C. pisiformis was confirmed to have the potential to fight inflammation in LPS-Mediated RAW264.7 murine macrophages. CCK8 test showed that rCpStefin below 50 μg/mL concentration did not affect cellular viability. Moreover, the NO production level determined by the Griess test was decreased. In addition, the secretion levels of IL-1β, IL-6, and TNF-α as measured by ELISA were decreased. Furthermore, it exerted anti-inflammatory activity by decreasing the production of proinflammatory cytokines and proinflammatory mediators, including IL-1β, IL-6, TNF-α, iNOS, and COX-2 at the gene transcription level, as measured by qRT-PCR. Therefore, Type I cystatin derived from C. pisiformis suppresses the LPS-Mediated inflammatory response of the intermediate host and is a potential candidate for the treatment of inflammatory diseases.
Full article
(This article belongs to the Section Parasitology)
►▼
Show Figures
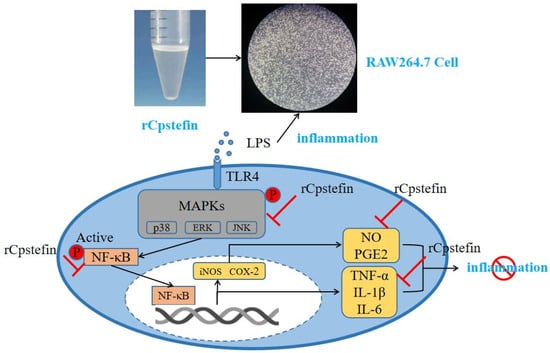
Figure 1
Open AccessArticle
Prosthetic Joint Infections Caused by Mycobacterium tuberculosis Complex—An ESGIAI–ESGMYC Multicenter, Retrospective Study and Literature Review
by
Alvaro Auñon, Llanos Salar-Vidal, Ignacio Mahillo-Fernandez, Francisco Almeida, Pedro Pereira, Jaime Lora-Tamayo, Tristan Ferry, Sarah Souèges, Aurélien Dinh, Rosa Escudero, Candela Menéndez Fernández-Miranda, Alicia Rico, Nicolo Rossi and Jaime Esteban
Microorganisms 2024, 12(5), 849; https://doi.org/10.3390/microorganisms12050849 - 24 Apr 2024
Abstract
Purpose: While tuberculosis remains a significant global health concern, prosthetic joint infections (PJIs) caused by members of the Mycobacterium tuberculosis complex are exceptionally rare. Our objective is to perform a retrospective search of new cases of this disease and analyze all cases available
[...] Read more.
Purpose: While tuberculosis remains a significant global health concern, prosthetic joint infections (PJIs) caused by members of the Mycobacterium tuberculosis complex are exceptionally rare. Our objective is to perform a retrospective search of new cases of this disease and analyze all cases available in the literature of tuberculous PJIs, aiming to detect factors that may influence patient outcomes. Methods: The ESGIAI and ESGMYC study groups were used to collect information on non-published cases of tuberculous prosthetic joint infections (PJIs). Additionally, a literature review of all published cases of tuberculous PJIs was conducted. All identified cases in the retrospective study and in the literature review were merged and included in the statistical analysis, involving both univariate and multivariate analyses. Results: Fifteen previously unreported cases of tuberculous prosthetic joint infections (PJIs) from four countries were detailed. Among them, ten patients were female, with a median age of 76 years. The hip was affected in 13 cases. Seven patients experienced co-infection with another microorganism. Treatment approaches varied, with 13 patients undergoing implant removal, one treated with DAIR (debridement, antibiotics, and implant retention), and one case was treated with an unknown treatment method. All patients received antibiotic therapy and achieved a cure. The literature review that was conducted detected 155 published cases. Univariate analysis revealed a statistical significance for previous tuberculosis, joint, and no importance of surgery for cure. Conclusions: Tuberculous prosthetic joint infection (PJI) is a rare condition, typically presenting as a localized chronic infection. Antibiotic treatment is essential for the management of these patients, but neither surgical treatment nor duration of treatment seems to have importance in the outcome.
Full article
(This article belongs to the Special Issue Device-Related Infections and Bacterial Biofilms)
Open AccessArticle
Evaluation of the Increased Genetic Resolution and Utility for Source Tracking of a Recently Developed Method for Genotyping Cyclospora cayetanensis
by
Susan R. Leonard, Mark K. Mammel, Sonia Almeria, Solomon T. Gebru, David K. Jacobson, Anna C. Peterson, Joel L. N. Barratt and Steven M. Musser
Microorganisms 2024, 12(5), 848; https://doi.org/10.3390/microorganisms12050848 - 24 Apr 2024
Abstract
Cyclospora cayetanensis is a foodborne parasite that causes cyclosporiasis, an enteric illness in humans. Genotyping methods are used to genetically discriminate between specimens from cyclosporiasis cases and can complement source attribution investigations if the method is sufficiently sensitive for application to food items.
[...] Read more.
Cyclospora cayetanensis is a foodborne parasite that causes cyclosporiasis, an enteric illness in humans. Genotyping methods are used to genetically discriminate between specimens from cyclosporiasis cases and can complement source attribution investigations if the method is sufficiently sensitive for application to food items. A very sensitive targeted amplicon sequencing (TAS) assay for genotyping C. cayetanensis encompassing 52 loci was recently designed. In this study, we analyzed 66 genetically diverse clinical specimens to assess the change in phylogenetic resolution between the TAS assay and a currently employed eight-marker scheme. Of the 52 markers, ≥50 were successfully haplotyped for all specimens, and these results were used to generate a hierarchical cluster dendrogram. Using a previously described statistical approach to dissect hierarchical trees, the 66 specimens resolved into 24 and 27 distinct genetic clusters for the TAS and an 8-loci scheme, respectively. Although the specimen composition of 15 clusters was identical, there were substantial differences between the two dendrograms, highlighting the importance of both inclusion of additional genome coverage and choice of loci to target for genotyping. To evaluate the ability to genetically link contaminated food samples with clinical specimens, C. cayetanensis was genotyped from DNA extracted from raspberries inoculated with fecal specimens. The contaminated raspberry samples were assigned to clusters with the corresponding clinical specimen, demonstrating the utility of the TAS assay for traceback efforts.
Full article
(This article belongs to the Section Food Microbiology)
►▼
Show Figures
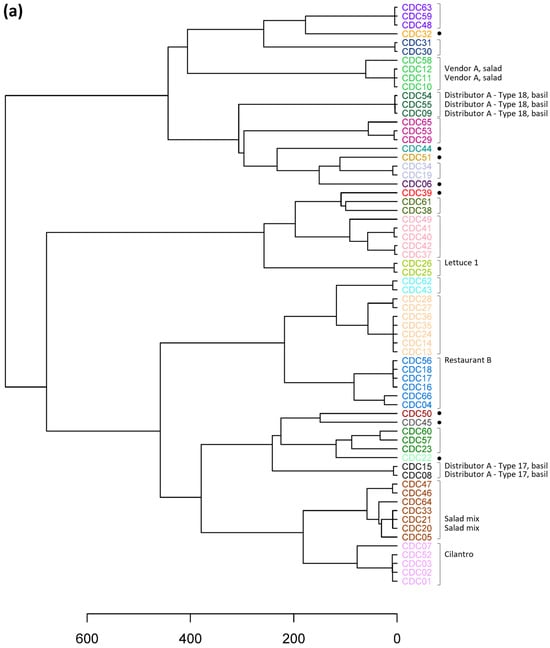
Figure 1
Open AccessSystematic Review
Evidence for Immunity against Tetanus, Diphtheria, and Pertussis through Natural Infection or Vaccination in Adult Solid Organ Transplant Recipients: A Systematic Review
by
Emil Lenzing, Zitta Barrella Harboe, Søren Schwartz Sørensen, Allan Rasmussen, Susanne Dam Nielsen and Omid Rezahosseini
Microorganisms 2024, 12(5), 847; https://doi.org/10.3390/microorganisms12050847 - 24 Apr 2024
Abstract
(1) Background: We aim to systematically review the current evidence on immunity against tetanus, diphtheria, and pertussis in adult solid organ transplantation (SOT) recipients, either through natural infection or vaccination. (2) Methods: This systematic review was conducted per PRISMA guidelines. We assessed the
[...] Read more.
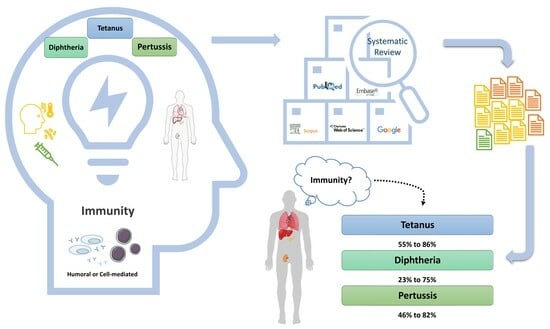
(1) Background: We aim to systematically review the current evidence on immunity against tetanus, diphtheria, and pertussis in adult solid organ transplantation (SOT) recipients, either through natural infection or vaccination. (2) Methods: This systematic review was conducted per PRISMA guidelines. We assessed the risk of bias using the Cochrane RoB 2 and ROBINS-I and summarized the findings narratively due to the heterogeneity of the studies. (3) Results: Of the 315 screened articles, 11 were included. Tetanus immunity varied between 55% and 86%, diphtheria immunity from 23% to 75%, and pertussis immunity was between 46% and 82%. Post-vaccination immunity showed variation across the studies, with some indicating reductions and others no change, with antibody responses influenced by transplanted organs, gender, age, and immunosuppressive regimens. The single randomized study exhibited a low risk of bias, while of the ten non-randomized studies, six showed moderate and four serious risks of bias, necessitating cautious interpretation of results. (4) Conclusions: SOT recipients exhibit considerable immunity against tetanus and diphtheria at transplantation, but this immunity decreases over time. Although vaccination can enhance this immunity, the response may be suboptimal, and the increased antibody levels may not persist, underscoring the need for tailored vaccination strategies in this vulnerable population.
Full article
(This article belongs to the Special Issue Infections in Solid Organ Transplant Recipients)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Surfing the Waves of SARS-CoV-2: Analysis of Viral Genome Variants Using an NGS Survey in Verona, Italy
by
Emil Tonon, Riccardo Cecchetto, Erica Diani, Nicoletta Medaina, Giona Turri, Anna Lagni, Virginia Lotti and Davide Gibellini
Microorganisms 2024, 12(5), 846; https://doi.org/10.3390/microorganisms12050846 - 24 Apr 2024
Abstract
The availability of new technologies for deep sequencing, including next-generation sequencing (NGS), allows for the detection of viral genome variations. The epidemiological determination of SARS-CoV-2 viral genome changes during the pandemic waves displayed the genome evolution and subsequent onset of variants over time.
[...] Read more.
The availability of new technologies for deep sequencing, including next-generation sequencing (NGS), allows for the detection of viral genome variations. The epidemiological determination of SARS-CoV-2 viral genome changes during the pandemic waves displayed the genome evolution and subsequent onset of variants over time. These variants were often associated with a different impact on viral transmission and disease severity. We investigated, in a retrospective study, the trend of SARS-CoV-2-positive samples collected from the start of the Italian pandemic (January 2020) to June 2023. In addition, viral RNAs extracted from 938 nasopharyngeal swab samples were analyzed using NGS between February 2022 and June 2023. Sequences were analyzed with bioinformatic tools to identify lineages and mutations and for phylogenetic studies. Six pandemic waves were detected. In our samples, we predominantly detected BA.2, BQ.1, BA.5.1, BA.5.2, and, more recently, XBB.1 and its subvariants. The data describe the SARS-CoV-2 genome evolution involved in viral interactions with the host and the dynamics of specific genome mutations and deletions.
Full article
(This article belongs to the Section Virology)
►▼
Show Figures
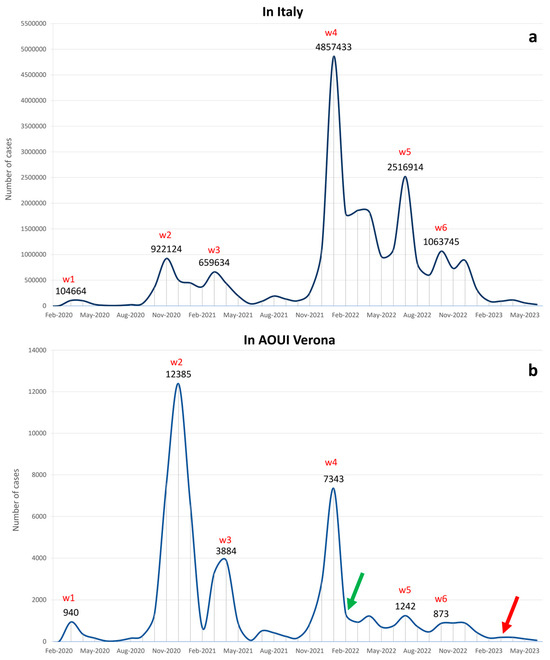
Figure 1
Open AccessArticle
Comparative Genomics Reveals Genetic Diversity and Variation in Metabolic Traits in Fructilactobacillus sanfranciscensis Strains
by
Xiaxia He, Yujuan Yu, Rober Kemperman, Luciana Jimenez, Faizan Ahmed Sadiq and Guohua Zhang
Microorganisms 2024, 12(5), 845; https://doi.org/10.3390/microorganisms12050845 - 23 Apr 2024
Abstract
Fructilactobacillus sanfranciscensis is a significant and dominant bacterial species of sourdough microbiota from ecological and functional perspectives. Despite the remarkable prevalence of different strains of this species in sourdoughs worldwide, the drivers behind the genetic diversity of this species needed to be clarified.
[...] Read more.
Fructilactobacillus sanfranciscensis is a significant and dominant bacterial species of sourdough microbiota from ecological and functional perspectives. Despite the remarkable prevalence of different strains of this species in sourdoughs worldwide, the drivers behind the genetic diversity of this species needed to be clarified. In this research, 14 F. sanfranciscensis strains were isolated from sourdough samples to evaluate the genetic diversity and variation in metabolic traits. These 14 and 31 other strains (obtained from the NCBI database) genomes were compared. The values for genome size and GC content, on average, turned out to 1.31 Mbp and 34.25%, respectively. In 45 F. sanfranciscensis strains, there were 162 core genes and 0 to 51 unique genes present in each strain. The primary functions of core genes were related to nucleotide, lipid transport, and amino acid, as well as carbohydrate metabolism. The size of core genes accounted for 41.18% of the pan-genome size in 14 F. sanfranciscensis strains, i.e., 0.70 Mbp of 1.70 Mbp. There were genetic variations among the 14 strains involved in carbohydrate utilization and antibiotic resistance. Moreover, exopolysaccharides biosynthesis-related genes were annotated, including epsABD, wxz, wzy. The Type IIA & IE CRISPR-Cas systems, pediocin PA-1 and Lacticin_3147_A1 bacteriocins operons were also discovered in F. sanfranciscensis. These findings can help to select desirable F. sanfranciscensis strains to develop standardized starter culture for sourdough fermentation, and expect to provide traditional fermented pasta with a higher quality and nutritional value for the consumers.
Full article
(This article belongs to the Special Issue Food Microorganisms and Genomics)
►▼
Show Figures
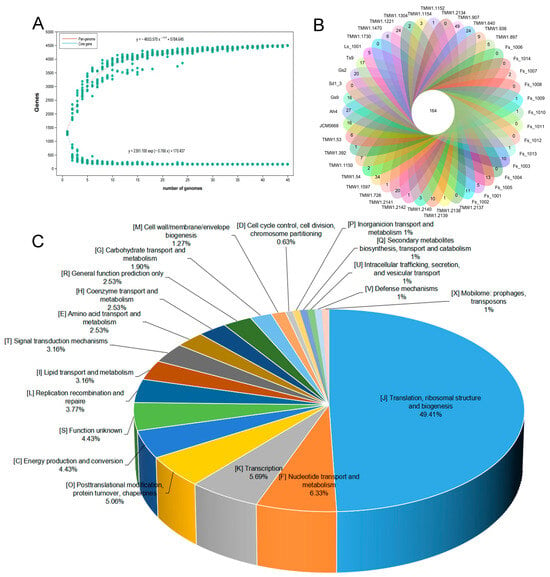
Figure 1
Open AccessArticle
Genomic Characterization and Molecular Detection of Rehmannia Allexivirus Virus, a Novel Allexivirus Infecting Rehmannia glutinosa
by
Yanhong Qin, Shuhao Lu, Yi Wen, Shaojian Li, Suxia Gao, Yuxia Liu, Xuemeng Li, Jin Yang, Fengli Wang, Fei Wang and Chuantao Lu
Microorganisms 2024, 12(5), 844; https://doi.org/10.3390/microorganisms12050844 - 23 Apr 2024
Abstract
Rehmannia glutinosa is one of the most important medicinal plants in China and is affected by viral diseases. In this study, a new virus tentatively named Rehmannia Allexivirus virus (ReAV) was identified through high-throughput sequencing, reverse-transcription polymerase chain reaction (RT-PCR), and Sanger sequencing.
[...] Read more.
Rehmannia glutinosa is one of the most important medicinal plants in China and is affected by viral diseases. In this study, a new virus tentatively named Rehmannia Allexivirus virus (ReAV) was identified through high-throughput sequencing, reverse-transcription polymerase chain reaction (RT-PCR), and Sanger sequencing. The complete genome length was 7297 nt and it contained five open reading frames (ORFs) encoding replicase, triple gene block 1(TGB1), TGB2, TGB3, and coat protein (CP). The replicase and CP presented nucleotide homology ranges of 59.9–65.2% and 47.5–55.5% between the nine ReAV isolates and the other 12 species of the genus Allexivirus. In the nine isolates, ReAV-20 and ReAV-31 isolates showed breakpoints in the replicase and CP regions, respectively. The other isolates shared 87.2–96.5% nt with the whole genome nucleotide identity. The phylogenetic tree showed that seven ReAV isolates based on replicase, CP, and whole genome sequences were clustered in the same branch and were related to the genus Allexivirus. The ReAV detection rates for 60 R. glutinosa samples were 73.3–81.7% through RT-PCR using primers targeting the replicase or CP genes. These results demonstrate that ReAV is the dominant virus in R. glutinosa. This study provides important evidence for understanding viruses infecting R. glutinosa and for establishing efficient strategies to prevent viral spread.
Full article
(This article belongs to the Section Microbial Biotechnology)
►▼
Show Figures
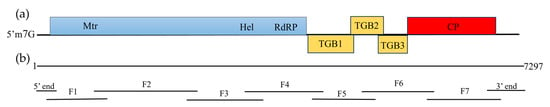
Figure 1
Open AccessArticle
Chemical Characterization and Effect of a Lactobacilli-Postbiotic on Streptococcus mutans Biofilm In Vitro
by
Guilherme Bandeira Santana, Patrick Veras Quelemes, Enedina Rodrigues da Silva Neta, Sidney Gonçalo de Lima and Gláuber Campos Vale
Microorganisms 2024, 12(5), 843; https://doi.org/10.3390/microorganisms12050843 - 23 Apr 2024
Abstract
Postbiotic is the term used to define the soluble factors, metabolic products, or byproducts released by live probiotic bacteria or after its lysis. The objective of this study was to carry out the chemical characterization of the postbiotic of Lacticaseibacillus rhamnosus LR-32 and
[...] Read more.
Postbiotic is the term used to define the soluble factors, metabolic products, or byproducts released by live probiotic bacteria or after its lysis. The objective of this study was to carry out the chemical characterization of the postbiotic of Lacticaseibacillus rhamnosus LR-32 and to evaluate its in vitro effect on the development of the Streptococcus mutans biofilm. After the cultivation of the probiotic strain, the postbiotic was extracted by centrifuging the culture and filtering the supernatant. This postbiotic was characterized by using gas chromatography coupled with mass spectrometry (GC–MS), and then it was used to determine the growth inhibition of S. mutans in its planktonic form; additionally, its effects on the following parameters in 48 h biofilm were evaluated: viable bacteria, dry weight, and gene expression of glucosyltransferases and VicR gene. The control group consisted of the biofilm without any treatment. A paired t-test was performed for statistical analysis, with the p-value set at 5%. Seventeen compounds of various chemical classes were identified in the postbiotic, including sugars, amino acids, vitamins, and acids. The treatment with the postbiotic led to an inhibition of the growth of S. mutans in its planktonic form, as well as a decrease in the number of viable bacteria, reduction in dry weight, and a negative regulation of the gene expression of gtfB, gtfC, gtfD, and vicR in its biofilm state, compared with the nontreated group (p < 0.05). The postbiotic of L. rhamnosus impaired the development of S. mutans biofilm.
Full article
(This article belongs to the Special Issue A Contemporary Look at Oral Microbe Management)
►▼
Show Figures
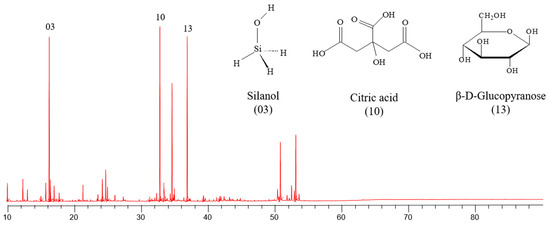
Figure 1
Open AccessReview
Tackling the Antimicrobial Resistance “Pandemic” with Machine Learning Tools: A Summary of Available Evidence
by
Doris Rusic, Marko Kumric, Ana Seselja Perisin, Dario Leskur, Josipa Bukic, Darko Modun, Marino Vilovic, Josip Vrdoljak, Dinko Martinovic, Marko Grahovac and Josko Bozic
Microorganisms 2024, 12(5), 842; https://doi.org/10.3390/microorganisms12050842 - 23 Apr 2024
Abstract
Antimicrobial resistance is recognised as one of the top threats healthcare is bound to face in the future. There have been various attempts to preserve the efficacy of existing antimicrobials, develop new and efficient antimicrobials, manage infections with multi-drug resistant strains, and improve
[...] Read more.
Antimicrobial resistance is recognised as one of the top threats healthcare is bound to face in the future. There have been various attempts to preserve the efficacy of existing antimicrobials, develop new and efficient antimicrobials, manage infections with multi-drug resistant strains, and improve patient outcomes, resulting in a growing mass of routinely available data, including electronic health records and microbiological information that can be employed to develop individualised antimicrobial stewardship. Machine learning methods have been developed to predict antimicrobial resistance from whole-genome sequencing data, forecast medication susceptibility, recognise epidemic patterns for surveillance purposes, or propose new antibacterial treatments and accelerate scientific discovery. Unfortunately, there is an evident gap between the number of machine learning applications in science and the effective implementation of these systems. This narrative review highlights some of the outstanding opportunities that machine learning offers when applied in research related to antimicrobial resistance. In the future, machine learning tools may prove to be superbugs’ kryptonite. This review aims to provide an overview of available publications to aid researchers that are looking to expand their work with new approaches and to acquaint them with the current application of machine learning techniques in this field.
Full article
(This article belongs to the Special Issue Latest Review Papers in Antimicrobial Agents and Resistance 2024)
►▼
Show Figures
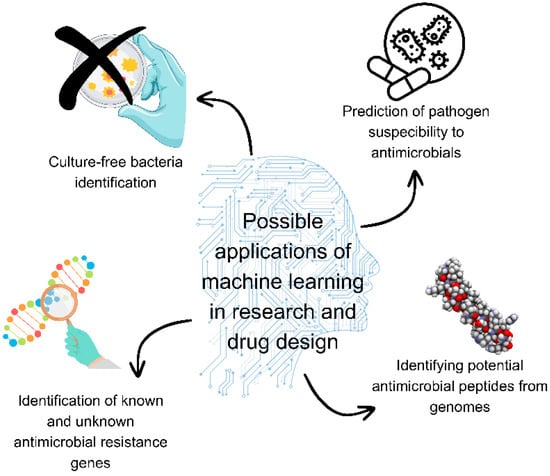
Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, Ecologies, Forests, Microorganisms, Plants
Litter Decompositions: From Individuals to Ecosystems
Topic Editors: Wen Zhou, Guihua LiuDeadline: 30 April 2024
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024
Topic in
Agronomy, Environments, Microorganisms, Pollutants, Sustainability, Water
Soil and Water Pollution Process and Remediation Technologies, 2nd Volume
Topic Editors: Hongbiao Cui, Ru Wang, Yu Shi, Haiying Lu, Lin ChenDeadline: 15 July 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Plant and Human Probiotics: Consequences on the Autochthonous Microbiota
Guest Editors: Elisa Gamalero, Elisa Bona, Giorgia Novello, Francesco VuoloDeadline: 30 April 2024
Special Issue in
Microorganisms
Human Skin Microbiota 2.0
Guest Editors: Holger Brüggemann, Rolf LoodDeadline: 15 May 2024
Special Issue in
Microorganisms
The Urban Microbiome
Guest Editors: Haruo Suzuki, Soojin JangDeadline: 31 May 2024
Special Issue in
Microorganisms
Emerging Pathogens Causing Acute Hepatitis
Guest Editors: Ibrahim M. Sayed, Sayed F. Abdelwahab, Ahmed El-ShamyDeadline: 15 June 2024
Topical Collections
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu